Sci-Kit Learn Simple Model Validation
Making predictions is easy, but getting them right is difficult. An essential part of predictive analytics is model validation - checking the quality of predictions. Those post looks at a simple model validation technique you can use with your machine learning models.
Once you have trained your predictive model it needs to make good predictions in the real world, with data it has not seen. There is a danger that this data does not look like the data used to train the model, and the model is unable to make accurate predictions. This could be because of the amount of noise in the training data, or fundamental differences between the training data and what occurs in the real world.
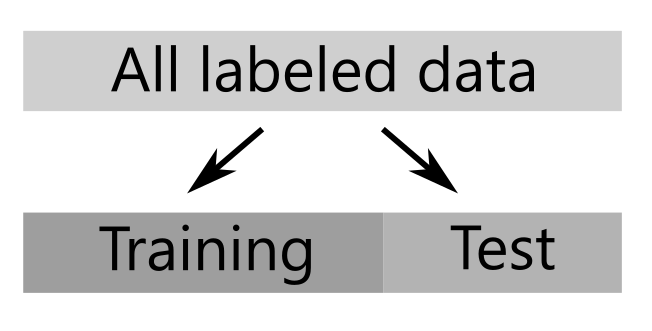
To get a more realistic view of model performance we need to some how test the model performance against ‘real world’ data. We can do this with a technique known as ‘cross-validation’. There are various cross-validation strategies you could adopt, but a good place to start is to simply separate your data into two sets. One set which we use to train to train the model, and one set which we keep back to use as a test of the model performance.
The test set is not used to train the model and can be used to give a better estimate of the model’s real world performance. The scikit-learn python package provides some useful helper functions for separating out a test set from the all the labelled data.
1
2
from sklearn.model_selection import train_test_split
X_train, x_test, y_train, y_test = train_test_split(X, y)
This gives you X_train and y_train : the data and outcomes you need to train your model. Once your model is trained on the training data, you can use the predict(X_test) method of the fitted model to get prediction results.
1
y_predict = clf.predict(X_test)
You can compare these predictions with the true y-values in y_test. Scikit-learn has performance metrics for evaluating classification and regression models.
A Simple Example
Here is a more complete example, which builds on my simple sklearn classification post.
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
from sklearn import datasets
from sklearn.ensemble import RandomForestClassifier
import pandas as pd
# Import sklearn train-test split
from sklearn.model_selection import train_test_split
# Import some sklearn classification metrics
from sklearn.metrics import accuracy_score, precision_score, recall_score, confusion_matrix, roc_auc_score
# Get sample dataset from sklearn datasets
cancer = datasets.load_breast_cancer()
X = cancer.data # (m,n) numpy array
y = cancer.target # (m,) numpy array
# Create training and testing data
# By default this will use 75% of the data for training
# and keep 25% for testing.
X_train, X_test, y_train, y_test = train_test_split(X, y)
# Train model with training data
clf_train = RandomForestClassifier()
clf_train.fit(X_train,y_train)
# Get predictions using training data and unseen test data
y_predict_test = clf_train.predict(X_test)
y_predict_train = clf_train.predict(X_train)
# Create table of metric scores
scores = {}
scores['accuracy'] = (accuracy_score(y_test, y_predict_test),
accuracy_score(y_train, y_predict_train))
scores['roc_auc'] = (roc_auc_score(y_test, y_predict_test),
roc_auc_score(y_train, y_predict_train))
scores['precision'] = (precision_score(y_test, y_predict_test),
precision_score(y_train, y_predict_train))
scores['recall'] = (recall_score(y_test, y_predict_test),
recall_score(y_train, y_predict_train))
scores_df = pd.DataFrame(scores).transpose()
scores_df.columns = ['Test', 'Train']
scores_df['Test - Train'] = scores_df.Test - scores_df.Train
print(scores_df)
| Test | Train | Test - Train | |
|---|---|---|---|
| accuracy | 0.944056 | 0.997653 | -0.053597 |
| precision | 0.945055 | 0.996283 | -0.051228 |
| recall | 0.966292 | 1.000000 | -0.033708 |
| roc_auc | 0.936850 | 0.996835 | -0.059986 |
From these scores you can see that the test score is worse than than the training score in all included metrics. This is to be expected as the model had not seen the test data. The test scores give a more realistic estimate of the real world performance of the model.