Organising Computational Projects
As computational projects grow the folder structure of scripts, configuration files and outputs can often become bloated and confused. You can help prevent this by spending some time early on to get the folder structure right. This is a useful guide to organising computational projects: https://www.ploscompbiol.org/article/info%3Adoi%2F10.1371%2Fjournal.pcbi.1000424
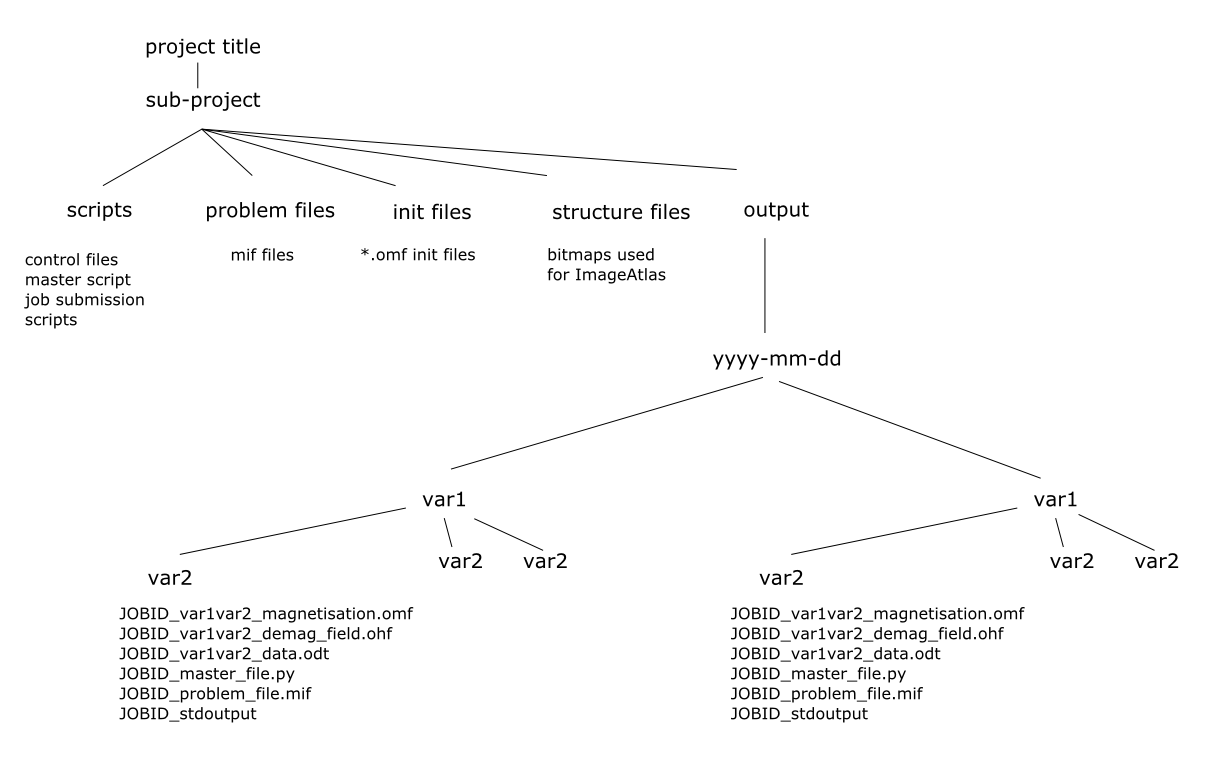
I often got lost trying to find “that simulation” from 6 months ago, so these suggestions were really good for me. After following those suggestions, my current directory tree is something like:
I’m sure this arrangement isn’t perfect. It has probably become a little bloated as the project has progressed, but I’m still sure that it is better than throwing all the files into one folder and hoping to sort it out later. I also have a master script which, as well as running project subscripts, also makes copies of them into the output folder (pre-pended with the job ID) so that I know exactly what was used to run a particular calculation. I would also emphasise the usefulness of having unique job IDs for calculations, with a record (digital or paper) of what those job IDs correspond to e.g. parameters, initialisation files etc. Combining job IDs with a search tool like Everything makes finding files relating to a particular calculation or simulation easy. I look in my lab book for a particular experiment/calculation/simulation and I can find all data relating to it with a simple search.